Hi GTDB curation team,
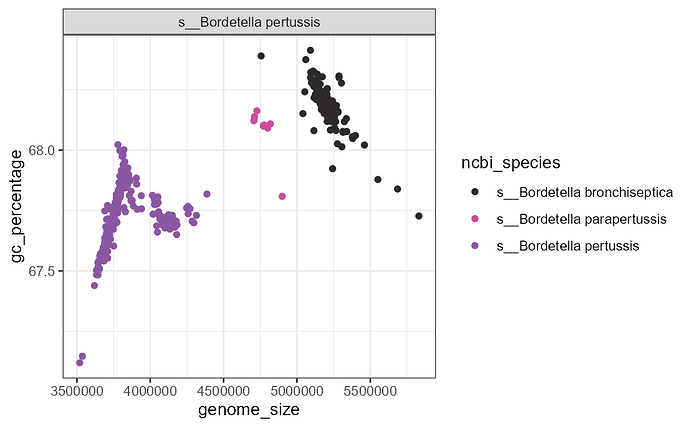
I have been going through some basic data in GTDB r226 and found s__Bordetella pertussisto display uneven genome_size and GC distribution. Moreover, I found all isolates over 4.7Mb to have the NCBI_taxonomy of either Bordetella parapertussis or Bordetella bronchiseptica, while all isolates under 4.3Mb to have NCBI_taxonomy assigned to Bordetella pertussis. It appears like parapertussis and bronchiseptica share ANI of over 99%, and ANIs between them and pertussis are both over 98%.
I am wondering if the large difference (~1.5Mb) in genome size justifies my suggestion of splitting
s__Bordetella pertussis into perhaps 2 or 3 seperated GTDB species.I am new to the GTDB community and may not be fully aware of the criterion for GTDB species curation process. Also, I am also not an expert in Bordetella. So I apologize if there are any mistakes.
Cheers,
David