Hello, I am trying to get a good bacterial phylogenetic tree that I can then annotate. The general GTDB bac_120 tree is too large for my needs and the GTDB-Tk v2 backbone tree seems like it will work much better (I do not need a high resolution tree). From what I’ve read, it looks like I’ll have to run the GTDB-Tk v2 workflow to generate this tree. Is there a way for me to obtain this tree file without downloading the entire program and running it?
Hi there, @nikhilgupta ![]()
Not sure if you’ve got something to suit your needs already, and I’m not sure how to get the gtdb-tk v2 backbone tree, but I’m writing in another avenue in case it helps.
I often want a relatively less dense tree spanning the current known diversity of a domain or phylum. I added functionality to my GToTree program a couple years ago for filtering down the GTDB for that very purpose.
One thing i commonly do is get all GTDB species representatives of the wanted group (80,789 bacterial at the time of doing this), randomly select 1 for each of the rank Order (1,653 currently), and then make a base reference tree from that (or in my case, usually add my new MAGs to them to make a new tree with all together).
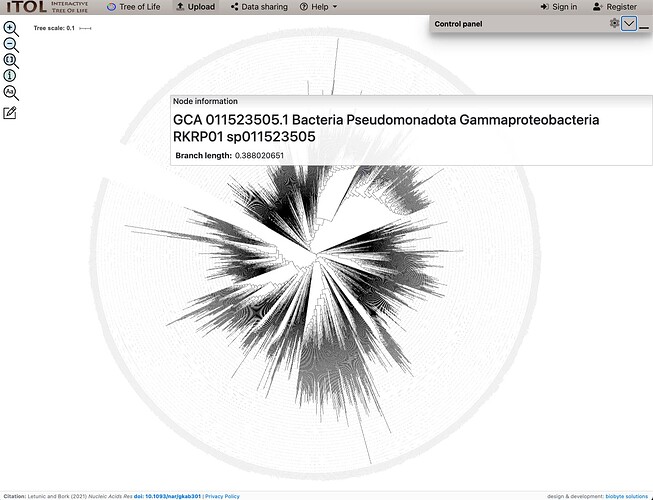
I ran one of these when I saw your message here, planning to attach the output tree and alignment file (in case you want to tree with something other than FastTree), but then realized only images are allowed to be attached. So here’s an image from IToL:
If you think this might help, the code is below to do it, or I can email you the alignment and tree file. It took about 2 hours as run below on a server using 20 cpus, and would probably take about 6 hours on a typical laptop.
Cheers!
mamba create -y -n gtotree -c astrobiomike -c conda-forge -c bioconda -c defaults gtotree==1.8.4
conda activate gtotree
### getting all bacterial accessions from GTDB
gtt-get-accessions-from-GTDB -t Bacteria --GTDB-representatives-only
#
# Reading in the GTDB info table...
# Using GTDB v214.1: Released Jun 9th, 2023
#
#
# The rank 'domain' has 394932 Bacteria entries.
#
# In considering only GTDB representative genomes:
#
# The rank 'domain' has 80789 Bacteria representative genome entries.
#
# The targeted NCBI accessions were written to:
# GTDB-Bacteria-domain-GTDB-rep-accs.txt
#
# A subset GTDB table of these targets was written to:
# GTDB-Bacteria-domain-GTDB-rep-metadata.tsv
### subsetting down to just 1 representative per Order
gtt-subset-GTDB-accessions -i GTDB-Bacteria-domain-GTDB-rep-metadata.tsv --get-only-individuals-for-the-rank order
#
# 80,789 initial entries were subset down to 1,653
#
# Subset accessions file for GToTree written to:
# subset-accessions.txt
#
# A subset GTDB taxonomy table for these accessions written to:
# subset-accessions-taxonomy.tsv
### then making the tree with GToTree
GToTree -a subset-accessions.txt -H Bacteria -D -j 20 -o bacterial-order
# where:
# -a is the list of accessions we want
# -H is specifying to used the pre-built Bacterial SGC-set HMMs (has 74 target genes)
# -D says to add GTDB taxonomy to the labels (by default: Domain, Phylum, Class, Species, Strain)